Data for tomorrow
Streamlining the journey from complex data to actionable insights
Speed up safe and sustainable product development by leveraging the power of AI, knowledge graphs and NLP
Gain access to the most recent and relevant data on disease genomics to accelerate drug R&D and unlock new precision medicine possibilities.
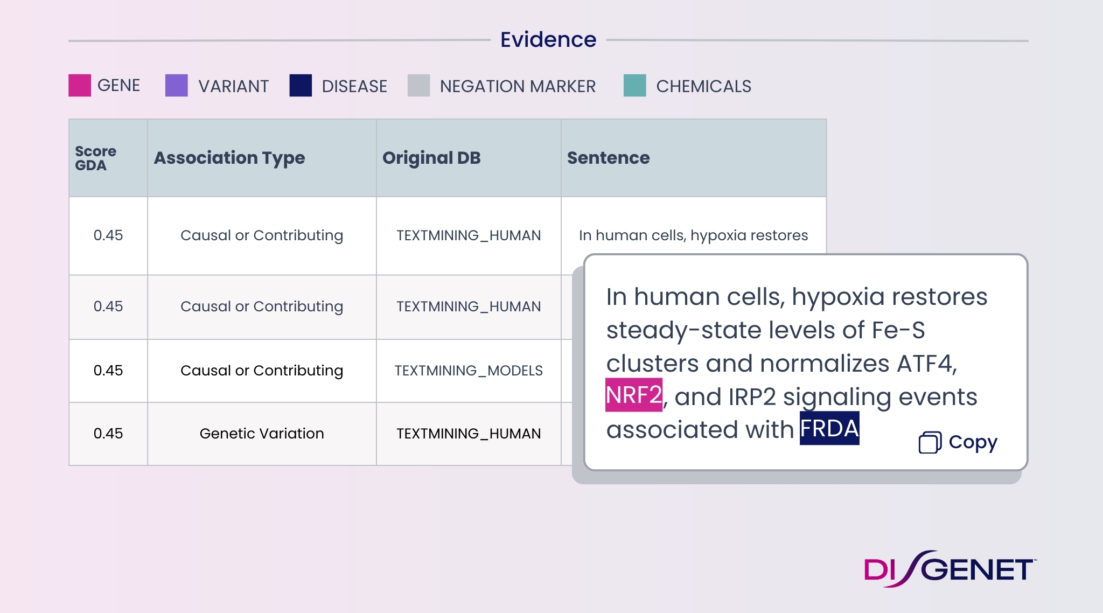
Natural Language Processing
Our state-of the-art NLP solutions make your textual data searchable, analyzable and actionable. We help you speed up informed innovation with data unlocked from text.
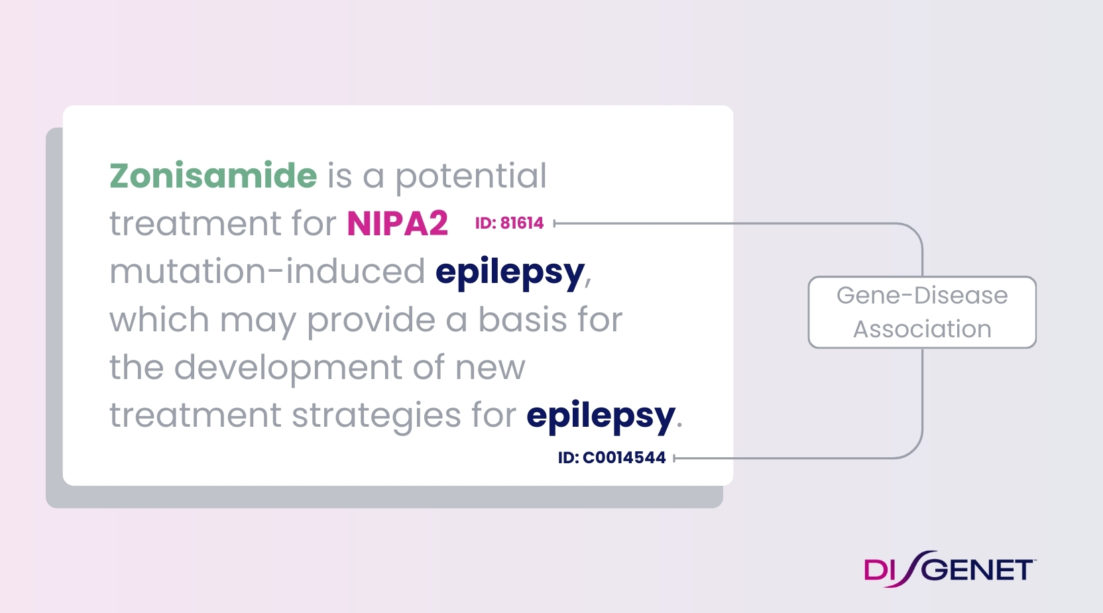
AI & Knowledge Graphs
We enable you to reveal insights from complex biological networks providing fine-grained, comprehensive coverage of intricate relationships between biomolecules.
DISGENET, the world’s most reliable & extensive gene-disease database
Get immediate access to information comparable to having read over 30 million articles.

50M+
disease associations
+7,200
citations worldwide
92%
NLP F-score
Est.
2010

MedBioinformatics, expanding DISGENET’s potential
MedBioinformatics is built on over 15 years of life science expertise. We unlock the true potential of your data through cutting-edge methods in data analytics, semantic integration, and NLP. This empowers you to develop innovative and safer products, prioritizing human health and well-being.